For researchers today, our current challenge lies in the need to enrich siloed and unstructured scientific data.
There are hundreds of publicly available databases containing information about the life sciences and healthcare. For example, PubMed Central® (PMC) is a free full-text archive of biomedical and life sciences journal literature at the U.S. National Institutes of Health's National Library of Medicine (NIH/NLM). Europe PMC is the European version of PMC.
There are specialized databases for almost every therapeutic area, and for some therapeutic areas, there are aggregators of public domain databases, such as the Cancer Data Research Commons.
But while there is an abundance of data, much of it is siloed and is available “as is,” meaning that there’s been little value added on top of these disparate, often unstructured databases.
Another way of putting this is that the databases are “unenriched.”
The Many Forms of Data Enrichment
Data enrichment can take many forms:
- Data can be classified and tagged to make searching easier and facilitate the clustering of results.
- Sentiment scores can be added to individual records.
- A variety of machine learning techniques can be used to identify the data that would be of most interest to a particular researcher. A data enrichment challenge is applying these techniques to disparate datasets from different sources in a consistent manner.
Applying Consistent, Structured Metadata
Foundation is a service aggregating about 20 databases, most in the public domain, unstructured, and related to life sciences. We enrich the content in these databases by adding a consistent layer of structured metadata to facilitate granular searching, information discovery, and uncovering hidden connections. Our proprietary, machine-learning-driven tagging engine generates the tags and categories assigned to each piece of content.
As the technological landscape evolves, we recognize the significance of staying at the forefront of data enrichment methods to support advanced scientific research effectively. It is crucial to continually enhance our understanding and implementation of these methods to better serve the evolving needs of the scientific community.
An early enhancement in this area was adding PubChem tagging to several of our key databases: PubMed, Patents, Clinical Trials, Grants, and Tech Transfer. This allowed more fine-tuned searches for documents mentioning a specific chemical compound and allowed a researcher to quickly get basic information about a compound mentioned in a document.
Customer response to this capability was highly positive, so we looked for more metadata to enrich our corpus.
We were aware of the Unified Medical Language System (UMLS) and the National Center for Biomedical Ontology (NCBO). The goal was to annotate our key datasets further and simultaneously build a process by which we could tag our customers' proprietary datasets with these more specific taxonomies.
Fast & Secure Data Enrichment at Scale
The NCBO hosts BioPortal, “the world’s most comprehensive repository of biomedical ontologies." To annotate a corpus with these ontologies, a tool called the OntoPortal Virtual Appliance has been developed and successfully implemented in several environments. Unfortunately, we tried to use the Appliance, but ran into many hurdles that would not be easy to overcome.
The two most critical issues were:
-
The Appliance didn’t scale; we wanted to tag many, many millions of documents, and the Appliance was not designed for this.
-
There were security vulnerabilities such that the OntoPortal Appliance would not be SOC 2 compliant, an essential requirement for ResoluteAI customers.
The BioPortal presented a treasure trove of data enrichment opportunities waiting to be accessed, but we needed to find or build a new way to deploy it.
We then looked into SciSpacy, an open-sourced library by the Allen Institute of AI containing spaCy models for processing biomedical, scientific, or clinical text. SciSpacy can link entities to UMLS concepts1, though not to specific ontologies within UMLS. The pipeline uses a neural network2 trained on several datasets, including MedMentions and Ontonotes.
SciSpacy can take as input a given text and output extracted entities and map those entities to concepts in UMLS. However, SciSpacy's actual entity linker (to UMLS) needed to be faster for our purposes due to the large number of records and entities we needed to match. Furthermore, we needed to link entities to various specific ontologies rather than just to higher-level concepts. Lastly, we could not rely on them updating their files as new revisions get released by UMLS. However, the NCBO tools were useful for seeing how they handled UMLS metafile parsing edge cases and their heuristics for choosing ontologies relevant to a text.
Although we did not use the NCBO BioPortal, nor most of the SciSpacy Entity Linker pipeline, looking at the methodology and implementation of these tools, as well as recognizing their shortcomings, drove the evolution of our ontology linker.
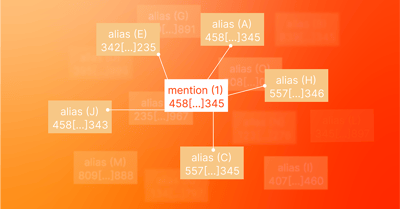
Our implementation uses SciSpacy's entity extraction models with our own UMLS parser and entity linker. The entity linker is much like SciSpacy's, except it is built to scale using Spark, a distributed data processing cluster. We make use of approximate nearest neighbor search3 in Spark between billions of mentions and millions of candidate aliases. Unlike SciSpacy, we are able to match on specific UMLS ontologies.
In all, Research Solutions' ResoluteAI solution is two orders of magnitude faster than the SciSpacy method.
Enriching Data with New Ontologies, Taxonomies, & Controlled VocabulariesDespite a number of fits and starts, we were ultimately able to annotate a number of large research databases with several ontologies4, taxonomies, and controlled vocabularies available through UMLS. This has dramatically improved our faceted search features, allowing users to filter search results using metadata combinations. Analytics, especially heatmaps, have also been improved. For large datasets such as PubMed and Patents, this accelerates the research process and, in several cases to date, has been extremely helpful in white space searches.
We are currently working on three next steps:
-
Enrichment of publicly available databases with additional hierarchical metadata has proven valuable to researchers. We plan to extend this capability to customer-proprietary datasets as part of our Nebula enterprise search platform. Most corporate research data is poorly tagged and haphazardly managed; internal research can be dramatically improved by adding more structured, domain-specific metadata.
-
We selected the taxonomies from UMLS that we believed would have the most immediate impact on our customers' workflow. We have already had requests to enrich our public domain databases further with more UMLS taxonomies and several from the Open Biological and Biomedical Ontology (OBO) Foundry.
-
While data enrichment has apparent benefits, it can contribute to information overload in some respects. To address this challenge, we incorporate many of these ontologies, taxonomies, and controlled vocabularies into a graph database, allowing users to view the connections between content from disparate databases more efficiently. By clustering search results around annotations or combinations of annotations, users will be able to quickly identify papers, patents, clinical trials, etc., that meet highly granular specifications.
Discover the Power & Potential of Enriched Data With Our Product Suite
Harness the capability to transform your research process, uncover new insights, and unlock hidden potential in your information assets. We invite you to explore a live demo of our comprehensive suite of products and experience firsthand the dramatic improvements in research productivity our solutions provide.
Schedule a demo today and propel your organization towards data-informed decision making.
1 SciSpacy's entity linker matches entities to UMLS concepts based on textual similarity. It works as follows: it extracts entities from the text called mentions. For each mention, it does a nearest neighbor search in a pre-computed index of UMLS concept aliases in the UMLS knowledge base. To do the nearest neighbors search (ANN), a mention of text is converted into a numerical vector of numbers based on a TF-IDF analysis of its 3-character n-grams. In other words, the text is broken down into n-grams like this: "biomedical" → [bio iom ome med edi dic ica al] and then each individual n-gram is assigned a weight indicating how "important" it is based on its frequency of occurrence within each concept alias vs. across all concepts. The final result is a vector that can be compared for similarity to other vectors. The similarity between the vectors indicates a substantial similarity between the texts they represent and is robust enough to ignore minor spelling differences and word inversions. This robustness is another advantage over the OntoPortal usage of mgrep since mrep cannot recognize that minor variations are likely equivalent.
2 A specialized entity extractor powered by a neural network trained on medical and scientific corpora performs entity extraction better than simpler methods such as word boundary tokenization. For example, it will recognize phrases like "Post Traumatic Stress Disease" as a single concept.
3 Approximate Nearest Neighbor Search is a proximity search for points (e.g., UMLS aliases) close to a given query point (e.g., extracted mentions). We use LSH (locality-sensitive hashing) but have begun transitioning to HNSW (hierarchical navigable small worlds).
4 MeSH, MedDRA, RxNorm, OMIM, GO, SNOMED CT, and ICD-10.